🌊 Random Forest: Deep Dive & Best Practices
Concise, clear, and validated revision notes on Random Forest with Python & Scikit-learn — practical best practices for beginners and practitioners.
Table of Contents
- Introduction
- Core Concepts
- Algorithm Workflow
- Terminology Tables
- Hyperparameters
- Feature Importance
- Out-of-Bag (OOB) Error
- Hyperparameter Tuning
- Complete Implementation Examples
- Advantages
- Limitations and When to Avoid
- Best Practices
- Common Pitfalls and Solutions
- Advanced Techniques
- Comparison with Other Algorithms
- Mathematical Foundation
- Real-World Applications
- Debugging and Troubleshooting
- Performance Optimization Checklist
- Summary
- References
Introduction
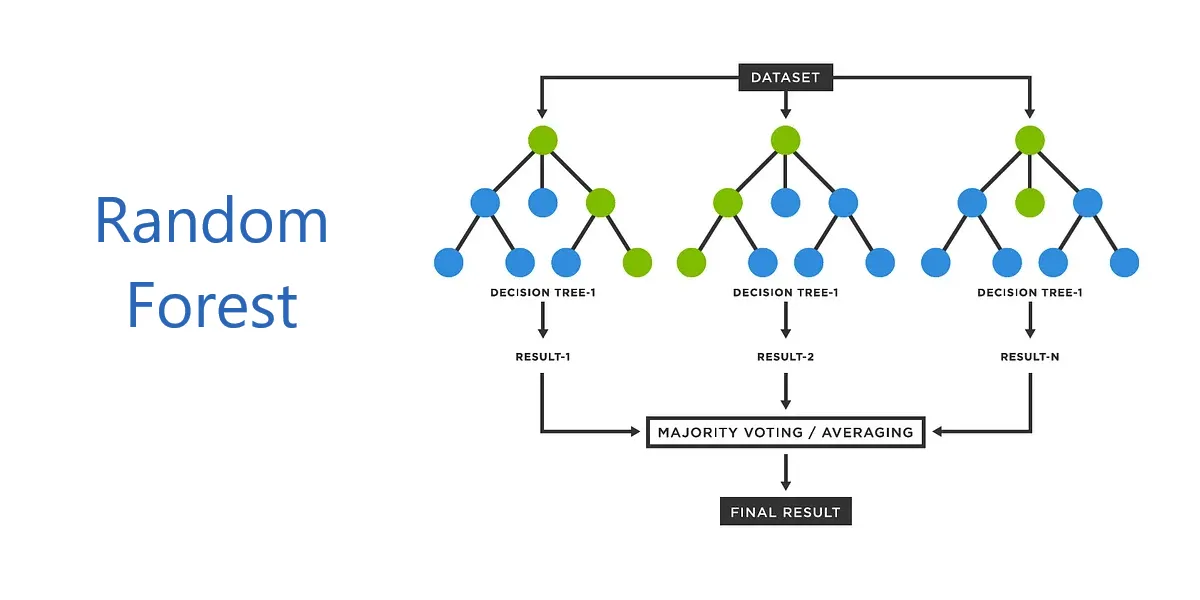
Random Forest is a powerful ensemble learning algorithm that combines multiple decision trees to produce accurate and robust predictions. Developed by Leo Breiman and Adele Cutler in 2001, it has become one of the most widely-used machine learning algorithms due to its simplicity, versatility, and strong performance across diverse applications.
Random Forest works by constructing numerous decision trees during training and outputting the mode of classes for classification tasks or the mean prediction for regression tasks. This ensemble approach significantly reduces overfitting and variance compared to individual decision trees.
Core Concepts
Ensemble Learning
Ensemble learning combines predictions from multiple models to achieve better performance than any single model. Random Forest employs two key ensemble techniques:
Bagging (Bootstrap Aggregating): Each tree is trained on a random sample (bootstrap sample) drawn with replacement from the original training data. Approximately 67% of the data is included in each bootstrap sample, leaving about 33% as out-of-bag samples for validation.
Feature Randomness: At each node split, only a random subset of features is considered for finding the best split, rather than evaluating all features. This decorrelates the trees and reduces variance.
Decision Trees Foundation
Before understanding Random Forests, it is essential to grasp decision trees. A decision tree is a supervised learning algorithm that recursively splits data based on feature values to minimize impurity at each node.
Splitting Criteria:
- Classification: Gini impurity or Information Gain (entropy)
- Regression: Variance reduction
Decision trees are prone to overfitting when grown to maximum depth, capturing noise and specific patterns from training data. Random Forests address this by averaging multiple deep trees.
Algorithm Workflow
Training Process
Step 1: Bootstrap Sampling
For each tree \(t\) in the forest (where \(t = 1, 2, ..., N\)):
- Draw \(n\) samples with replacement from the training set of size \(n\)
- This creates a bootstrap sample where approximately 63.2% are unique samples
- The remaining ~36.8% form the out-of-bag (OOB) set for that tree
Step 2: Random Feature Selection
At each node during tree construction:
- Randomly select \(m\) features from total \(p\) features (\(m < p\))
- Default values:
- Classification: \(m = \sqrt{p}\)
- Regression: \(m = p/3\)
- Find the best split among these \(m\) features only
Step 3: Tree Growing
- Grow each tree to maximum depth without pruning
- Each tree makes its own independent prediction
- No cross-tree communication during training
Step 4: Aggregation
For predictions:
- Classification: Majority voting across all trees
- Regression: Mean of predictions from all trees
Prediction Process
Given a new input \(x\):
Classification:
\[\hat{y} = \text{mode}\{h_1(x), h_2(x), ..., h_N(x)\}\]where \(h_i(x)\) is the prediction from tree \(i\)
Regression:
\[\hat{y} = \frac{1}{N}\sum_{i=1}^{N} h_i(x)\]Terminology Tables
Table 1: Process Stage Terminology
| General Term | Random Forest Specific | Ensemble Context | Statistical Context | Alternative Terms |
|---|---|---|---|---|
| Training | Forest Construction | Ensemble Building | Model Fitting | Learning Phase |
| Sampling | Bootstrap Sampling | Bagging | Resampling with Replacement | Data Aggregation |
| Splitting | Node Splitting | Feature Selection | Recursive Partitioning | Tree Growing |
| Aggregation | Voting/Averaging | Ensemble Combination | Prediction Pooling | Consensus Building |
| Validation | OOB Evaluation | Internal Validation | Holdout Testing | Cross-validation |
| Prediction | Inference | Ensemble Prediction | Forward Pass | Scoring |
Table 2: Component Hierarchy
| Level | Component | Description | Scope |
|---|---|---|---|
| 1 (Highest) | Random Forest | Complete ensemble model | Entire algorithm |
| 2 | Individual Trees | Decision tree estimators | Single classifier/regressor |
| 3 | Tree Structure | Nodes and branches | Tree architecture |
| 4 | Node | Split point | Single decision |
| 5 | Split | Feature threshold | Binary partition |
| 6 (Lowest) | Leaf | Terminal prediction | Final output node |
Table 3: Data Flow Terminology
| Stage | Input | Process | Output | Alternative Names |
|---|---|---|---|---|
| Initialization | Original Dataset | Bootstrap Sampling | Training Subsets | Data Preparation |
| Tree Construction | Bootstrap Sample | Recursive Splitting | Decision Tree | Model Building |
| Feature Selection | All Features | Random Sampling | Feature Subset | Attribute Sampling |
| Node Processing | Parent Node | Split Evaluation | Child Nodes | Branching |
| Aggregation | Tree Predictions | Voting/Averaging | Final Prediction | Ensemble Output |
Hyperparameters
Critical Hyperparameters
n_estimators
- Description: Number of trees in the forest
- Default: 100 (scikit-learn)
- Impact: More trees improve accuracy but increase computational cost
- Best Practice: Start with 100-500; increase until performance plateaus
- Range: 50-1000+ depending on dataset size
max_features
- Description: Number of features to consider for best split at each node
- Default:
- Classification:
'sqrt'(\(\sqrt{n_{features}}\)) - Regression:
'log2'or1.0(all features)
- Classification:
- Options:
'sqrt','log2',None, integer, or float - Impact: Controls tree correlation and randomness
- Best Practice: Use defaults; tune only if overfitting occurs
max_depth
- Description: Maximum depth of each tree
- Default:
None(nodes expand until pure or min_samples_split reached) - Impact: Deeper trees capture complex patterns but may overfit
- Best Practice: Start with
None; limit if overfitting or high memory usage
min_samples_split
- Description: Minimum samples required to split an internal node
- Default: 2
- Impact: Higher values prevent overfitting to small groups
- Best Practice: 2-20; increase for noisy data
min_samples_leaf
- Description: Minimum samples required at a leaf node
- Default: 1
- Impact: Smooths predictions; prevents overly specific leaves
- Best Practice: 1-10; higher for smoother decision boundaries
bootstrap
- Description: Whether to use bootstrap sampling
- Default:
True - Impact: Enables bagging and OOB evaluation
- Best Practice: Keep
Truefor standard random forest
oob_score
- Description: Whether to calculate out-of-bag score
- Default:
False - Impact: Provides internal validation metric
- Best Practice: Set
Truefor quick model assessment
Supporting Hyperparameters
random_state
- Description: Controls randomness for reproducibility
- Default:
None - Best Practice: Set to fixed integer (e.g., 42) for reproducible results
n_jobs
- Description: Number of parallel jobs
- Default:
None(single processor) - Options: -1 (all processors), 1, or specific number
- Best Practice: Use -1 for large datasets to leverage multicore processing
criterion
- Description: Split quality measure
- Default:
- Classification:
'gini' - Regression:
'squared_error'
- Classification:
- Options:
- Classification:
'gini','entropy','log_loss' - Regression:
'squared_error','absolute_error','friedman_mse','poisson'
- Classification:
- Best Practice: Defaults work well; experiment if domain-specific
Feature Importance
Random Forest provides built-in feature importance measures to identify which features contribute most to predictions.
Methods
1. Mean Decrease in Impurity (MDI)
Also called Gini importance. Calculates total reduction in node impurity weighted by probability of reaching that node across all trees.
Computation: For feature \(j\):
\[\text{Importance}(j) = \frac{1}{N}\sum_{t=1}^{N}\sum_{k \in \text{tree}_t} p_k \Delta i(k, j)\]where:
- \(N\) = number of trees
- \(p_k\) = proportion of samples reaching node \(k\)
- \(\Delta i(k, j)\) = decrease in impurity at node \(k\) using feature \(j\)
Advantages:
- Fast computation (calculated during training)
- Built into scikit-learn via
feature_importances_attribute
Limitations:
- Biased toward high-cardinality features (many unique values)
- Can overestimate importance of continuous features
- Based on training data only
2. Permutation Feature Importance
Measures decrease in model performance when a feature’s values are randomly shuffled.
Algorithm:
- Calculate baseline performance on validation set
- For each feature:
- Randomly shuffle feature values
- Calculate new performance
- Importance = baseline performance - shuffled performance
- Repeat multiple times and average
Advantages:
- Model-agnostic (works with any model)
- Not biased toward high-cardinality features
- Uses holdout/validation data
Limitations:
- Computationally expensive
- Slower than MDI
3. SHAP Values
SHAP (SHapley Additive exPlanations) provides both global and local feature importance based on game theory.
Advantages:
- Consistent and theoretically sound
- Provides local explanations for individual predictions
- Handles feature interactions
Limitations:
- Computationally intensive
- Requires external library (
shap)
Python Implementation
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
import numpy as np
import pandas as pd
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.inspection import permutation_importance
import matplotlib.pyplot as plt
# Load data
from sklearn.datasets import load_iris
X, y = load_iris(return_X_y=True)
feature_names = load_iris().feature_names
# Split data
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.3, random_state=42
)
# Train model
rf = RandomForestClassifier(n_estimators=100, random_state=42)
rf.fit(X_train, y_train)
# Method 1: MDI (Gini Importance)
importances_mdi = rf.feature_importances_
std = np.std([tree.feature_importances_ for tree in rf.estimators_], axis=0)
# Create DataFrame for visualization
importance_df = pd.DataFrame({
'Feature': feature_names,
'Importance': importances_mdi,
'Std': std
}).sort_values('Importance', ascending=False)
print("Mean Decrease in Impurity:")
print(importance_df)
# Plot MDI
plt.figure(figsize=(10, 6))
plt.barh(importance_df['Feature'], importance_df['Importance'])
plt.xlabel('Feature Importance')
plt.title('Feature Importance (Mean Decrease in Impurity)')
plt.tight_layout()
plt.show()
# Method 2: Permutation Importance
perm_importance = permutation_importance(
rf, X_test, y_test, n_repeats=10, random_state=42, n_jobs=-1
)
perm_df = pd.DataFrame({
'Feature': feature_names,
'Importance': perm_importance.importances_mean,
'Std': perm_importance.importances_std
}).sort_values('Importance', ascending=False)
print("\nPermutation Importance:")
print(perm_df)
# Plot Permutation Importance
plt.figure(figsize=(10, 6))
plt.barh(perm_df['Feature'], perm_df['Importance'])
plt.xlabel('Permutation Importance')
plt.title('Feature Importance (Permutation Method)')
plt.tight_layout()
plt.show()
Out-of-Bag (OOB) Error
Out-of-Bag error provides an internal validation mechanism unique to bagging-based algorithms like Random Forest.
Concept
When training with bootstrap sampling, approximately 36.8% of samples are left out for each tree. These out-of-bag (OOB) samples act as a validation set for that specific tree.
OOB Score Calculation:
- For each sample \(i\) in training set:
- Identify trees where sample \(i\) was out-of-bag
- Aggregate predictions from these trees only
- Compare with true label
- Calculate accuracy across all samples:
Advantages
- Free validation: No need for separate validation set
- Efficient: Computed during training without extra cost
- Unbiased estimate: Approximates cross-validation error
- Maximizes data usage: All data used for training
Limitations
- Only available when
bootstrap=True - May overestimate error in certain conditions:
- Small sample sizes
- Balanced class distributions
- High number of features relative to samples
Python Implementation
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import make_classification
import matplotlib.pyplot as plt
# Generate synthetic data
X, y = make_classification(
n_samples=1000,
n_features=20,
n_informative=15,
n_redundant=5,
random_state=42
)
# Track OOB error as trees are added
oob_errors = []
n_trees_range = range(10, 201, 10)
for n_trees in n_trees_range:
rf = RandomForestClassifier(
n_estimators=n_trees,
oob_score=True,
random_state=42,
n_jobs=-1
)
rf.fit(X, y)
oob_error = 1 - rf.oob_score_
oob_errors.append(oob_error)
print(f"Trees: {n_trees}, OOB Error: {oob_error:.4f}")
# Plot OOB error convergence
plt.figure(figsize=(10, 6))
plt.plot(n_trees_range, oob_errors, marker='o')
plt.xlabel('Number of Trees')
plt.ylabel('OOB Error Rate')
plt.title('Out-of-Bag Error vs Number of Trees')
plt.grid(True)
plt.tight_layout()
plt.show()
Comparison with Cross-Validation:
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
from sklearn.model_selection import cross_val_score
# OOB Score
rf_oob = RandomForestClassifier(
n_estimators=100,
oob_score=True,
random_state=42
)
rf_oob.fit(X, y)
print(f"OOB Score: {rf_oob.oob_score_:.4f}")
# Cross-Validation Score
rf_cv = RandomForestClassifier(n_estimators=100, random_state=42)
cv_scores = cross_val_score(rf_cv, X, y, cv=5)
print(f"CV Score: {cv_scores.mean():.4f} ± {cv_scores.std():.4f}")
Hyperparameter Tuning
Tuning Strategies
1. Grid Search
Exhaustively searches through a manually specified subset of hyperparameter space.
Advantages:
- Comprehensive search within specified range
- Guaranteed to find best combination in grid
Disadvantages:
- Computationally expensive
- Inefficient for large hyperparameter spaces
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import RandomForestClassifier
# Define parameter grid
param_grid = {
'n_estimators': [100, 200, 300],
'max_depth': [10, 20, 30, None],
'min_samples_split': [2, 5, 10],
'min_samples_leaf': [1, 2, 4],
'max_features': ['sqrt', 'log2']
}
# Initialize Random Forest
rf = RandomForestClassifier(random_state=42)
# Grid search with cross-validation
grid_search = GridSearchCV(
estimator=rf,
param_grid=param_grid,
cv=5,
n_jobs=-1,
verbose=2,
scoring='accuracy'
)
# Fit grid search
grid_search.fit(X_train, y_train)
# Best parameters
print("Best Parameters:", grid_search.best_params_)
print("Best CV Score:", grid_search.best_score_)
# Best model
best_rf = grid_search.best_estimator_
2. Randomized Search
Samples random combinations from hyperparameter distributions.
Advantages:
- More efficient than grid search
- Can explore wider range
- Good for initial exploration
Disadvantages:
- May miss optimal combination
- Requires more iterations for convergence
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import randint, uniform
# Define parameter distributions
param_distributions = {
'n_estimators': randint(100, 500),
'max_depth': [10, 20, 30, None],
'min_samples_split': randint(2, 20),
'min_samples_leaf': randint(1, 10),
'max_features': ['sqrt', 'log2', None],
'bootstrap': [True, False]
}
# Initialize Random Forest
rf = RandomForestClassifier(random_state=42)
# Randomized search
random_search = RandomizedSearchCV(
estimator=rf,
param_distributions=param_distributions,
n_iter=100, # number of random combinations
cv=5,
n_jobs=-1,
verbose=2,
random_state=42,
scoring='accuracy'
)
# Fit randomized search
random_search.fit(X_train, y_train)
# Results
print("Best Parameters:", random_search.best_params_)
print("Best CV Score:", random_search.best_score_)
3. Bayesian Optimization
Uses probabilistic model to guide search toward promising regions.
Advantages:
- Most efficient for expensive evaluations
- Learns from previous iterations
- Better than random search with fewer iterations
Disadvantages:
- Requires additional library
- More complex to implement
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
from skopt import BayesSearchCV
from skopt.space import Real, Categorical, Integer
# Define search space
search_spaces = {
'n_estimators': Integer(100, 500),
'max_depth': Integer(10, 50),
'min_samples_split': Integer(2, 20),
'min_samples_leaf': Integer(1, 10),
'max_features': Categorical(['sqrt', 'log2']),
}
# Initialize Bayesian optimization
bayes_search = BayesSearchCV(
estimator=RandomForestClassifier(random_state=42),
search_spaces=search_spaces,
n_iter=50,
cv=5,
n_jobs=-1,
verbose=2,
random_state=42
)
# Fit Bayesian search
bayes_search.fit(X_train, y_train)
print("Best Parameters:", bayes_search.best_params_)
print("Best CV Score:", bayes_search.best_score_)
Best Practices for Tuning
- Start with defaults: Random Forest works well with default parameters
- Prioritize tuning order:
- First:
n_estimators,max_features - Second:
max_depth,min_samples_split,min_samples_leaf - Last: Other parameters
- First:
- Use appropriate CV folds: 5-fold for most cases; 10-fold for small datasets
- Balance accuracy vs computational cost: More trees and deeper trees = slower
- Monitor overfitting: Check training vs validation scores
- Use OOB score for quick validation: Faster than CV during exploration
Complete Implementation Examples
Classification Example
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
import numpy as np
import pandas as pd
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import (
classification_report,
confusion_matrix,
roc_auc_score,
roc_curve
)
import matplotlib.pyplot as plt
import seaborn as sns
# Load data
data = load_breast_cancer()
X, y = data.data, data.target
feature_names = data.feature_names
# Split data
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=42, stratify=y
)
print(f"Training samples: {X_train.shape[0]}")
print(f"Testing samples: {X_test.shape[0]}")
print(f"Number of features: {X_train.shape[1]}")
# Initialize Random Forest Classifier
rf_clf = RandomForestClassifier(
n_estimators=200,
max_depth=20,
min_samples_split=5,
min_samples_leaf=2,
max_features='sqrt',
bootstrap=True,
oob_score=True,
random_state=42,
n_jobs=-1,
verbose=1
)
# Train model
print("\nTraining Random Forest Classifier...")
rf_clf.fit(X_train, y_train)
# OOB Score
print(f"\nOOB Score: {rf_clf.oob_score_:.4f}")
# Cross-validation
cv_scores = cross_val_score(
rf_clf, X_train, y_train, cv=5, scoring='accuracy'
)
print(f"CV Accuracy: {cv_scores.mean():.4f} ± {cv_scores.std():.4f}")
# Predictions
y_pred = rf_clf.predict(X_test)
y_pred_proba = rf_clf.predict_proba(X_test)[:, 1]
# Evaluation
print("\nClassification Report:")
print(classification_report(y_test, y_pred, target_names=data.target_names))
# Confusion Matrix
cm = confusion_matrix(y_test, y_pred)
plt.figure(figsize=(8, 6))
sns.heatmap(cm, annot=True, fmt='d', cmap='Blues',
xticklabels=data.target_names,
yticklabels=data.target_names)
plt.title('Confusion Matrix')
plt.ylabel('True Label')
plt.xlabel('Predicted Label')
plt.tight_layout()
plt.show()
# ROC Curve
fpr, tpr, thresholds = roc_curve(y_test, y_pred_proba)
roc_auc = roc_auc_score(y_test, y_pred_proba)
plt.figure(figsize=(8, 6))
plt.plot(fpr, tpr, color='darkorange', lw=2,
label=f'ROC curve (AUC = {roc_auc:.2f})')
plt.plot([0, 1], [0, 1], color='navy', lw=2, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver Operating Characteristic (ROC) Curve')
plt.legend(loc="lower right")
plt.grid(alpha=0.3)
plt.tight_layout()
plt.show()
# Feature Importance
importances = rf_clf.feature_importances_
indices = np.argsort(importances)[::-1][:10] # Top 10 features
plt.figure(figsize=(12, 6))
plt.title('Top 10 Feature Importances')
plt.bar(range(10), importances[indices])
plt.xticks(range(10), [feature_names[i] for i in indices], rotation=45, ha='right')
plt.ylabel('Importance')
plt.tight_layout()
plt.show()
Regression Example
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
33
34
35
36
37
38
39
40
41
42
43
44
45
46
47
48
49
50
51
52
53
54
55
56
57
58
59
60
61
62
63
64
65
66
67
68
69
70
71
72
73
74
75
76
77
78
79
80
81
82
83
84
85
86
87
88
89
90
91
92
93
94
95
96
97
98
99
100
101
102
103
104
import numpy as np
import pandas as pd
from sklearn.datasets import load_diabetes
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import (
mean_squared_error,
mean_absolute_error,
r2_score
)
import matplotlib.pyplot as plt
# Load data
data = load_diabetes()
X, y = data.data, data.target
feature_names = data.feature_names
# Split data
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=42
)
print(f"Training samples: {X_train.shape[0]}")
print(f"Testing samples: {X_test.shape[0]}")
# Initialize Random Forest Regressor
rf_reg = RandomForestRegressor(
n_estimators=200,
max_depth=15,
min_samples_split=5,
min_samples_leaf=2,
max_features='sqrt',
bootstrap=True,
oob_score=True,
random_state=42,
n_jobs=-1,
verbose=1
)
# Train model
print("\nTraining Random Forest Regressor...")
rf_reg.fit(X_train, y_train)
# OOB Score
print(f"\nOOB Score (R²): {rf_reg.oob_score_:.4f}")
# Cross-validation
cv_scores = cross_val_score(
rf_reg, X_train, y_train, cv=5, scoring='r2'
)
print(f"CV R² Score: {cv_scores.mean():.4f} ± {cv_scores.std():.4f}")
# Predictions
y_pred = rf_reg.predict(X_test)
# Evaluation
mse = mean_squared_error(y_test, y_pred)
rmse = np.sqrt(mse)
mae = mean_absolute_error(y_test, y_pred)
r2 = r2_score(y_test, y_pred)
print(f"\nTest Set Performance:")
print(f"RMSE: {rmse:.2f}")
print(f"MAE: {mae:.2f}")
print(f"R² Score: {r2:.4f}")
# Prediction vs Actual Plot
plt.figure(figsize=(10, 6))
plt.scatter(y_test, y_pred, alpha=0.6, edgecolors='k')
plt.plot([y_test.min(), y_test.max()],
[y_test.min(), y_test.max()],
'r--', lw=2)
plt.xlabel('Actual Values')
plt.ylabel('Predicted Values')
plt.title('Random Forest Regression: Predicted vs Actual')
plt.grid(alpha=0.3)
plt.tight_layout()
plt.show()
# Residuals Plot
residuals = y_test - y_pred
plt.figure(figsize=(10, 6))
plt.scatter(y_pred, residuals, alpha=0.6, edgecolors='k')
plt.axhline(y=0, color='r', linestyle='--', lw=2)
plt.xlabel('Predicted Values')
plt.ylabel('Residuals')
plt.title('Residual Plot')
plt.grid(alpha=0.3)
plt.tight_layout()
plt.show()
# Feature Importance
importances = rf_reg.feature_importances_
indices = np.argsort(importances)[::-1]
plt.figure(figsize=(10, 6))
plt.title('Feature Importances')
plt.bar(range(len(importances)), importances[indices])
plt.xticks(range(len(importances)),
[feature_names[i] for i in indices],
rotation=45, ha='right')
plt.ylabel('Importance')
plt.tight_layout()
plt.show()
Advantages
- High Accuracy: Ensemble approach produces superior predictions compared to individual trees
- Robust to Overfitting: Averaging multiple trees reduces variance and prevents overfitting
- Handles Missing Data: Can maintain performance with missing values through surrogate splits
- No Feature Scaling Required: Tree-based nature makes it invariant to feature scaling
- Feature Importance: Provides interpretable feature importance scores
- Versatility: Works for both classification and regression tasks
- Non-parametric: Makes no assumptions about data distribution
- Parallel Processing: Trees can be trained independently, enabling multicore computation
- Built-in Validation: OOB error provides unbiased performance estimate
- Resistant to Outliers: Ensemble voting reduces impact of extreme values
Limitations and When to Avoid
Limitations
- Computational Cost: Training many deep trees is memory and time-intensive
- Prediction Speed: Slower inference compared to simpler models; evaluating all trees takes time
- Model Interpretability: Black-box nature makes it harder to explain than single decision trees
- Memory Requirements: Storing entire forest requires significant memory
- Extrapolation Issues: Cannot predict values outside training range in regression tasks
- Linear Relationships: May underperform on purely linear data compared to linear models
- High-Cardinality Bias: Feature importance can be biased toward features with many unique values
- Not Incremental: Cannot update model with new data; requires complete retraining
When NOT to Use Random Forest
1. Real-time Prediction Requirements
- Low-latency applications where millisecond response times are critical
- Consider: Linear models, shallow decision trees, or optimized neural networks
2. Limited Computational Resources
- Memory-constrained devices or embedded systems
- Consider: Logistic regression, Naive Bayes, or single decision tree
3. Linear Relationships
- Data with clear linear patterns
- Consider: Linear regression, Ridge, Lasso
4. High Interpretability Required
- Regulatory compliance requiring explainable decisions
- Medical diagnosis where decision paths must be transparent
- Consider: Single decision tree, linear models, or rule-based systems
5. Online Learning Needed
- Streaming data requiring incremental updates
- Consider: Online gradient descent, incremental decision trees
6. Time Series Forecasting
- Temporal dependencies requiring sequential modeling
- Cannot extrapolate trends beyond training data
- Consider: ARIMA, Prophet, LSTM, or specialized time series models
7. Very Small Datasets
- Fewer than 100-200 samples
- Risk of overfitting despite ensemble approach
- Consider: Regularized linear models, k-NN
8. Extremely Large Datasets
- Datasets with millions of samples and features
- Training time becomes prohibitive
- Consider: Gradient boosting (XGBoost, LightGBM), or neural networks with mini-batch training
Best Practices
Data Preparation
1. Handle Missing Values
1
2
3
4
5
6
7
8
9
from sklearn.impute import SimpleImputer
# For numerical features
imputer_num = SimpleImputer(strategy='median')
X_num = imputer_num.fit_transform(X_numerical)
# For categorical features
imputer_cat = SimpleImputer(strategy='most_frequent')
X_cat = imputer_cat.fit_transform(X_categorical)
2. No Feature Scaling Needed
- Random Forest is invariant to feature scaling
- Skip normalization/standardization for efficiency
3. Encode Categorical Variables
1
2
3
4
5
6
7
8
9
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
# Binary categorical: LabelEncoder
le = LabelEncoder()
X['binary_feature'] = le.fit_transform(X['binary_feature'])
# Multi-class categorical: OneHotEncoder
ohe = OneHotEncoder(sparse_output=False, handle_unknown='ignore')
X_encoded = ohe.fit_transform(X[['categorical_feature']])
4. Balance Classes (for Classification)
1
2
3
4
5
6
7
8
9
from imblearn.over_sampling import SMOTE
from imblearn.under_sampling import RandomUnderSampler
# For imbalanced datasets
smote = SMOTE(random_state=42)
X_balanced, y_balanced = smote.fit_resample(X_train, y_train)
# Or use class_weight parameter
rf = RandomForestClassifier(class_weight='balanced')
Model Training
1. Start with Defaults
1
2
3
4
5
rf = RandomForestClassifier(
n_estimators=100,
random_state=42,
n_jobs=-1
)
2. Monitor Training
1
2
3
# Track training progress
rf = RandomForestClassifier(verbose=1)
rf.fit(X_train, y_train)
3. Use Stratified Splitting (Classification)
1
2
3
4
5
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, stratify=y, random_state=42
)
4. Enable OOB Scoring
1
2
3
4
5
6
rf = RandomForestClassifier(
oob_score=True,
bootstrap=True
)
rf.fit(X_train, y_train)
print(f"OOB Score: {rf.oob_score_}")
Model Evaluation
1. Multiple Metrics
1
2
3
4
5
6
7
8
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
y_pred = rf.predict(X_test)
print(f"Accuracy: {accuracy_score(y_test, y_pred):.4f}")
print(f"Precision: {precision_score(y_test, y_pred, average='weighted'):.4f}")
print(f"Recall: {recall_score(y_test, y_pred, average='weighted'):.4f}")
print(f"F1 Score: {f1_score(y_test, y_pred, average='weighted'):.4f}")
2. Cross-Validation
1
2
3
4
5
6
7
8
9
10
11
from sklearn.model_selection import cross_validate
# Multiple scoring metrics
scoring = ['accuracy', 'precision_weighted', 'recall_weighted', 'f1_weighted']
scores = cross_validate(
rf, X, y, cv=5, scoring=scoring, n_jobs=-1
)
for metric in scoring:
print(f"{metric}: {scores[f'test_{metric}'].mean():.4f} ± {scores[f'test_{metric}'].std():.4f}")
3. Learning Curves
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
from sklearn.model_selection import learning_curve
train_sizes, train_scores, val_scores = learning_curve(
rf, X, y, cv=5, n_jobs=-1,
train_sizes=np.linspace(0.1, 1.0, 10),
scoring='accuracy'
)
# Plot learning curves
plt.figure(figsize=(10, 6))
plt.plot(train_sizes, train_scores.mean(axis=1), label='Training score')
plt.plot(train_sizes, val_scores.mean(axis=1), label='Validation score')
plt.xlabel('Training Set Size')
plt.ylabel('Score')
plt.legend()
plt.title('Learning Curves')
plt.grid(alpha=0.3)
plt.tight_layout()
plt.show()
Optimization
1. Incremental Tuning
1
2
3
4
5
6
7
8
9
10
11
# Step 1: Tune n_estimators
for n in [50, 100, 200, 300]:
rf = RandomForestClassifier(n_estimators=n, random_state=42)
scores = cross_val_score(rf, X_train, y_train, cv=5)
print(f"n_estimators={n}: {scores.mean():.4f}")
# Step 2: Tune max_features
for mf in ['sqrt', 'log2', None]:
rf = RandomForestClassifier(n_estimators=200, max_features=mf, random_state=42)
scores = cross_val_score(rf, X_train, y_train, cv=5)
print(f"max_features={mf}: {scores.mean():.4f}")
2. Early Stopping (Monitor Performance)
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
# Track performance as trees are added
estimators_range = range(10, 201, 10)
train_scores = []
test_scores = []
for n_est in estimators_range:
rf = RandomForestClassifier(n_estimators=n_est, random_state=42)
rf.fit(X_train, y_train)
train_scores.append(rf.score(X_train, y_train))
test_scores.append(rf.score(X_test, y_test))
# Plot to identify plateau
plt.figure(figsize=(10, 6))
plt.plot(estimators_range, train_scores, label='Train')
plt.plot(estimators_range, test_scores, label='Test')
plt.xlabel('Number of Estimators')
plt.ylabel('Accuracy')
plt.legend()
plt.title('Score vs Number of Trees')
plt.grid(alpha=0.3)
plt.tight_layout()
plt.show()
3. Feature Selection
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
from sklearn.feature_selection import SelectFromModel
# Train initial model
rf = RandomForestClassifier(n_estimators=100, random_state=42)
rf.fit(X_train, y_train)
# Select features based on importance threshold
selector = SelectFromModel(rf, threshold='median', prefit=True)
X_train_selected = selector.transform(X_train)
X_test_selected = selector.transform(X_test)
print(f"Original features: {X_train.shape[1]}")
print(f"Selected features: {X_train_selected.shape[1]}")
# Retrain with selected features
rf_selected = RandomForestClassifier(n_estimators=100, random_state=42)
rf_selected.fit(X_train_selected, y_train)
print(f"Accuracy with selected features: {rf_selected.score(X_test_selected, y_test):.4f}")
Production Deployment
1. Model Serialization
1
2
3
4
5
6
7
8
9
10
11
import joblib
import pickle
# Save model (joblib recommended for sklearn)
joblib.dump(rf, 'random_forest_model.pkl')
# Load model
loaded_rf = joblib.load('random_forest_model.pkl')
# Verify loaded model
print(f"Test accuracy: {loaded_rf.score(X_test, y_test):.4f}")
2. Prediction Optimization
1
2
3
4
5
6
7
# Batch predictions for efficiency
batch_predictions = rf.predict(X_test)
# Parallel predictions
rf = RandomForestClassifier(n_jobs=-1) # Use all CPU cores
rf.fit(X_train, y_train)
predictions = rf.predict(X_test)
3. Memory Management
1
2
3
4
5
6
7
8
9
10
11
12
13
14
# For large forests, consider reducing memory footprint
rf = RandomForestClassifier(
n_estimators=100,
max_depth=10, # Limit tree depth
min_samples_leaf=5 # Prune small leaves
)
# Use warm_start for incremental training
rf = RandomForestClassifier(warm_start=True, n_estimators=50)
rf.fit(X_train, y_train)
# Add more trees later
rf.n_estimators = 100
rf.fit(X_train, y_train) # Trains additional 50 trees
4. Model Monitoring
1
2
3
4
5
6
7
8
9
10
# Track predictions and confidence
predictions = rf.predict(X_new)
probabilities = rf.predict_proba(X_new)
# Log low-confidence predictions
low_confidence_mask = probabilities.max(axis=1) < 0.7
print(f"Low confidence predictions: {low_confidence_mask.sum()}")
# Flag for human review
review_needed = X_new[low_confidence_mask]
Common Pitfalls and Solutions
Pitfall 1: Too Few Trees
Problem: High variance in predictions Solution: Increase n_estimators to 100-300
1
2
3
4
5
# Bad
rf = RandomForestClassifier(n_estimators=10)
# Good
rf = RandomForestClassifier(n_estimators=200)
Pitfall 2: Deep Overfitting
Problem: Perfect training accuracy but poor test performance Solution: Limit tree depth and increase minimum samples
1
2
3
4
5
6
7
8
9
10
11
# Check for overfitting
rf.fit(X_train, y_train)
print(f"Train accuracy: {rf.score(X_train, y_train):.4f}")
print(f"Test accuracy: {rf.score(X_test, y_test):.4f}")
# If gap > 0.1, reduce overfitting
rf = RandomForestClassifier(
max_depth=15,
min_samples_split=10,
min_samples_leaf=5
)
Pitfall 3: Ignoring Class Imbalance
Problem: Model biased toward majority class Solution: Use class_weight='balanced' or resampling
1
2
3
4
5
6
7
8
9
10
# Check class distribution
print(pd.Series(y_train).value_counts())
# Address imbalance
rf = RandomForestClassifier(class_weight='balanced')
# Or use resampling
from imblearn.over_sampling import SMOTE
smote = SMOTE()
X_balanced, y_balanced = smote.fit_resample(X_train, y_train)
Pitfall 4: Not Using OOB Score
Problem: Wasting time on separate validation Solution: Enable oob_score=True for quick validation
1
2
3
4
5
6
rf = RandomForestClassifier(
oob_score=True,
bootstrap=True
)
rf.fit(X_train, y_train)
print(f"OOB Score: {rf.oob_score_:.4f}")
Pitfall 5: Inappropriate Feature Preprocessing
Problem: Unnecessary scaling or transformation Solution: Skip scaling; handle encoding properly
1
2
3
4
5
6
7
# Don't do this (unnecessary)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_scaled = scaler.fit_transform(X)
# Random Forest doesn't need scaling
rf.fit(X, y) # Use original features directly
Pitfall 6: Treating as Online Learner
Problem: Trying to incrementally update model Solution: Use warm_start or retrain completely
1
2
3
4
5
6
7
8
9
10
# Incremental approach (limited)
rf = RandomForestClassifier(warm_start=True, n_estimators=50)
rf.fit(X_train_batch1, y_train_batch1)
rf.n_estimators += 50
rf.fit(X_train_batch2, y_train_batch2) # Adds 50 more trees
# Or retrain with all data
rf = RandomForestClassifier(n_estimators=100)
rf.fit(X_all, y_all)
Pitfall 7: Misinterpreting Feature Importance
Problem: Over-relying on Gini importance Solution: Use permutation importance for unbiased estimates
1
2
3
4
5
6
7
8
9
from sklearn.inspection import permutation_importance
# Gini importance (can be biased)
gini_importance = rf.feature_importances_
# Permutation importance (more reliable)
perm_importance = permutation_importance(
rf, X_test, y_test, n_repeats=10, random_state=42
)
Advanced Techniques
1. Handling Imbalanced Data
1
2
3
4
5
6
7
8
9
10
from imblearn.ensemble import BalancedRandomForestClassifier
# Balanced Random Forest
brf = BalancedRandomForestClassifier(
n_estimators=100,
sampling_strategy='auto',
replacement=True,
random_state=42
)
brf.fit(X_train, y_train)
2. Probability Calibration
1
2
3
4
5
6
7
8
9
10
11
from sklearn.calibration import CalibratedClassifierCV
# Calibrate probabilities
calibrated_rf = CalibratedClassifierCV(
rf, method='sigmoid', cv=5
)
calibrated_rf.fit(X_train, y_train)
# Compare probabilities
raw_proba = rf.predict_proba(X_test)
calibrated_proba = calibrated_rf.predict_proba(X_test)
3. Partial Dependence Plots
1
2
3
4
5
6
7
8
9
10
from sklearn.inspection import PartialDependenceDisplay
# Visualize feature effects
features = [0, 1, (0, 1)] # Individual and interaction
fig, ax = plt.subplots(figsize=(12, 4))
PartialDependenceDisplay.from_estimator(
rf, X_train, features, ax=ax
)
plt.tight_layout()
plt.show()
4. Stacking with Random Forest
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
from sklearn.ensemble import StackingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
# Base models
estimators = [
('rf', RandomForestClassifier(n_estimators=100, random_state=42)),
('svm', SVC(probability=True, random_state=42))
]
# Stack with logistic regression
stacking = StackingClassifier(
estimators=estimators,
final_estimator=LogisticRegression(),
cv=5
)
stacking.fit(X_train, y_train)
5. Multi-output Random Forest
1
2
3
4
5
6
7
8
9
10
11
12
13
14
from sklearn.datasets import make_multilabel_classification
from sklearn.ensemble import RandomForestClassifier
from sklearn.multioutput import MultiOutputClassifier
# Generate multi-label data
X, y = make_multilabel_classification(
n_samples=1000, n_features=20, n_classes=3, random_state=42
)
# Multi-output classifier
multi_rf = MultiOutputClassifier(
RandomForestClassifier(n_estimators=100, random_state=42)
)
multi_rf.fit(X_train, y_train)
Comparison with Other Algorithms
Random Forest vs Gradient Boosting
| Aspect | Random Forest | Gradient Boosting |
|---|---|---|
| Training | Parallel (independent trees) | Sequential (each tree corrects previous) |
| Speed | Faster training | Slower training |
| Overfitting | More resistant | More prone (requires careful tuning) |
| Accuracy | Good | Often better with tuning |
| Interpretability | Moderate | Moderate |
| Hyperparameter Sensitivity | Less sensitive | Highly sensitive |
| Best For | General-purpose, noisy data | Competition, clean data |
Random Forest vs Decision Tree
| Aspect | Random Forest | Decision Tree |
|---|---|---|
| Accuracy | Higher (ensemble) | Lower (single model) |
| Overfitting | Less prone | Very prone |
| Interpretability | Harder to interpret | Easy to visualize |
| Training Time | Slower | Faster |
| Prediction Time | Slower | Faster |
| Variance | Low | High |
| Bias | Low | Can be high or low |
Random Forest vs Neural Networks
| Aspect | Random Forest | Neural Networks |
|---|---|---|
| Feature Engineering | Minimal required | Can learn features |
| Data Requirements | Works with small data | Needs large datasets |
| Training Time | Moderate | Slow (deep networks) |
| Hyperparameter Tuning | Simpler | Complex |
| Interpretability | Feature importance available | Black box |
| Structured Data | Excellent | Good |
| Unstructured Data | Not suitable | Excellent |
Mathematical Foundation
Gini Impurity
For a node with $K$ classes and class proportions $p_1, p_2, …, p_K$:
$\text{Gini}(t) = 1 - \sum_{i=1}^{K} p_i^2$
Measures impurity of node $t$. Lower values indicate purer nodes.
Entropy (Information Gain)
$\text{Entropy}(t) = -\sum_{i=1}^{K} p_i \log_2(p_i)$
Alternative splitting criterion. Information gain maximizes reduction in entropy.
Variance Reduction (Regression)
For regression, split quality measured by variance reduction:
$\Delta \text{Var} = \text{Var}(\text{parent}) - \left( \frac{n_{\text{left}}}{n} \text{Var}(\text{left}) + \frac{n_{\text{right}}}{n} \text{Var}(\text{right}) \right)$
Bootstrap Sample Probability
Probability that a sample is selected in bootstrap:
$P(\text{selected}) = 1 - \left(1 - \frac{1}{n}\right)^n \approx 1 - e^{-1} \approx 0.632$
Probability of being out-of-bag:
$P(\text{OOB}) \approx 1 - 0.632 = 0.368$
Feature Importance Formula
For feature $j$:
$\text{Importance}(j) = \frac{1}{N_{\text{trees}}}\sum_{t=1}^{N_{\text{trees}}} \sum_{k \in \text{nodes}_t} p(k) \cdot \Delta i(k, j) \cdot \mathbb{1}(v(k) = j)$
where:
- $p(k)$ = proportion of samples reaching node $k$
- $\Delta i(k, j)$ = impurity decrease at node $k$ using feature $j$
- $v(k)$ = feature used for split at node $k$
- $\mathbb{1}(\cdot)$ = indicator function
Real-World Applications
1. Healthcare
- Disease Prediction: Predicting diabetes, heart disease based on patient features
- Patient Stratification: Identifying high-risk patients for intervention
- Drug Response: Predicting patient response to treatment
2. Finance
- Credit Scoring: Assessing loan default risk
- Fraud Detection: Identifying fraudulent transactions
- Stock Price Movement: Predicting market trends
- Customer Churn: Identifying customers likely to leave
3. E-commerce
- Recommendation Systems: Product recommendations
- Price Optimization: Dynamic pricing strategies
- Demand Forecasting: Inventory management
- Customer Segmentation: Targeted marketing
4. Agriculture
- Crop Yield Prediction: Estimating harvest quantities
- Disease Detection: Identifying plant diseases from images
- Soil Quality Assessment: Predicting soil fertility
5. Environmental Science
- Species Classification: Identifying animal/plant species
- Climate Modeling: Weather and climate predictions
- Pollution Monitoring: Air quality forecasting
6. Manufacturing
- Predictive Maintenance: Equipment failure prediction
- Quality Control: Defect detection
- Process Optimization: Improving production efficiency
Debugging and Troubleshooting
Problem: Poor Performance
Diagnosis:
1
2
3
4
5
6
# Check baseline
from sklearn.dummy import DummyClassifier
dummy = DummyClassifier(strategy='most_frequent')
dummy.fit(X_train, y_train)
print(f"Baseline accuracy: {dummy.score(X_test, y_test):.4f}")
print(f"RF accuracy: {rf.score(X_test, y_test):.4f}")
Solutions:
- Increase
n_estimators - Tune
max_features - Check for data leakage
- Verify feature engineering
- Ensure proper train-test split
Problem: Slow Training
Diagnosis:
1
2
3
4
5
6
import time
start = time.time()
rf.fit(X_train, y_train)
duration = time.time() - start
print(f"Training time: {duration:.2f} seconds")
Solutions:
1
2
3
4
5
6
7
8
9
10
11
12
13
# Reduce complexity
rf = RandomForestClassifier(
n_estimators=100, # Reduce trees
max_depth=15, # Limit depth
max_samples=0.8, # Use subset of samples
n_jobs=-1 # Parallel processing
)
# Or use max_samples for faster training
rf = RandomForestClassifier(
max_samples=0.7, # Use 70% of samples per tree
n_jobs=-1
)
Problem: High Memory Usage
Solutions:
1
2
3
4
5
6
7
8
9
10
11
12
# Limit tree complexity
rf = RandomForestClassifier(
max_depth=10,
min_samples_leaf=10,
max_leaf_nodes=50
)
# Process in chunks
from sklearn.model_selection import learning_curve
train_sizes, train_scores, val_scores = learning_curve(
rf, X, y, train_sizes=np.linspace(0.1, 1.0, 5)
)
Problem: Inconsistent Results
Solution: Set random seed
1
2
3
# Ensure reproducibility
rf = RandomForestClassifier(random_state=42)
np.random.seed(42)
Performance Optimization Checklist
Training Phase
- Use
n_jobs=-1for parallel processing - Enable
warm_startfor incremental training - Set appropriate
max_samplesto reduce training time - Limit
max_depthif memory-constrained - Use
oob_score=Trueinstead of separate validation
Prediction Phase
- Batch predictions instead of single samples
- Use probability thresholds instead of hard predictions when possible
- Cache model predictions for repeated queries
- Consider model compression for deployment
Memory Management
- Limit tree depth and leaf nodes
- Use
max_leaf_nodesparameter - Prune features with low importance
- Store model in compressed format
Code Quality
- Set
random_statefor reproducibility - Use cross-validation for robust evaluation
- Monitor OOB score during development
- Log hyperparameters and results
- Version control trained models
Summary
Random Forest is a versatile and powerful ensemble learning algorithm suitable for diverse machine learning tasks. Its robustness to overfitting, built-in feature importance, and minimal hyperparameter tuning requirements make it an excellent choice for both beginners and practitioners.
Key Takeaways:
- Random Forest combines multiple decision trees through bagging and feature randomness
- Works well with default parameters; tuning improves performance marginally
- Provides interpretable feature importance
- OOB error offers free validation during training
- Resistant to overfitting compared to single decision trees
- Best for tabular data with complex, non-linear relationships
- Not ideal for real-time applications, online learning, or time series with trends
When to Choose Random Forest:
- Tabular structured data
- Both classification and regression tasks
- Need for feature importance
- Moderate dataset sizes (hundreds to hundreds of thousands of samples)
- Acceptable training time (not real-time)
- Non-linear relationships
Implementation Workflow:
- Start with default hyperparameters
- Enable OOB scoring for quick validation
- Check feature importance
- Tune hyperparameters if needed (n_estimators, max_features first)
- Use cross-validation for final evaluation
- Monitor performance on holdout test set
Random Forest remains one of the most reliable and widely-used algorithms in machine learning, balancing accuracy, interpretability, and ease of use.
References
Scikit-learn: Ensemble Methods - Random Forests
Scikit-learn: RandomForestClassifier API
Scikit-learn: RandomForestRegressor API
Breiman, L. (2001). Random Forests. Machine Learning, 45(1), 5-32
Biau, G., & Scornet, E. (2016). A Random Forest Guided Tour
Interpretable Machine Learning: Random Forest
Scikit-learn: Feature Importances Example